1) LLM-supported phage generation
Investigate whether genome-scale language modeling can help generate viable candidate phages with lytic characteristics.
Deep learning for enhanced phage therapy
An integrated computational and experimental approach to improve phage therapy design.
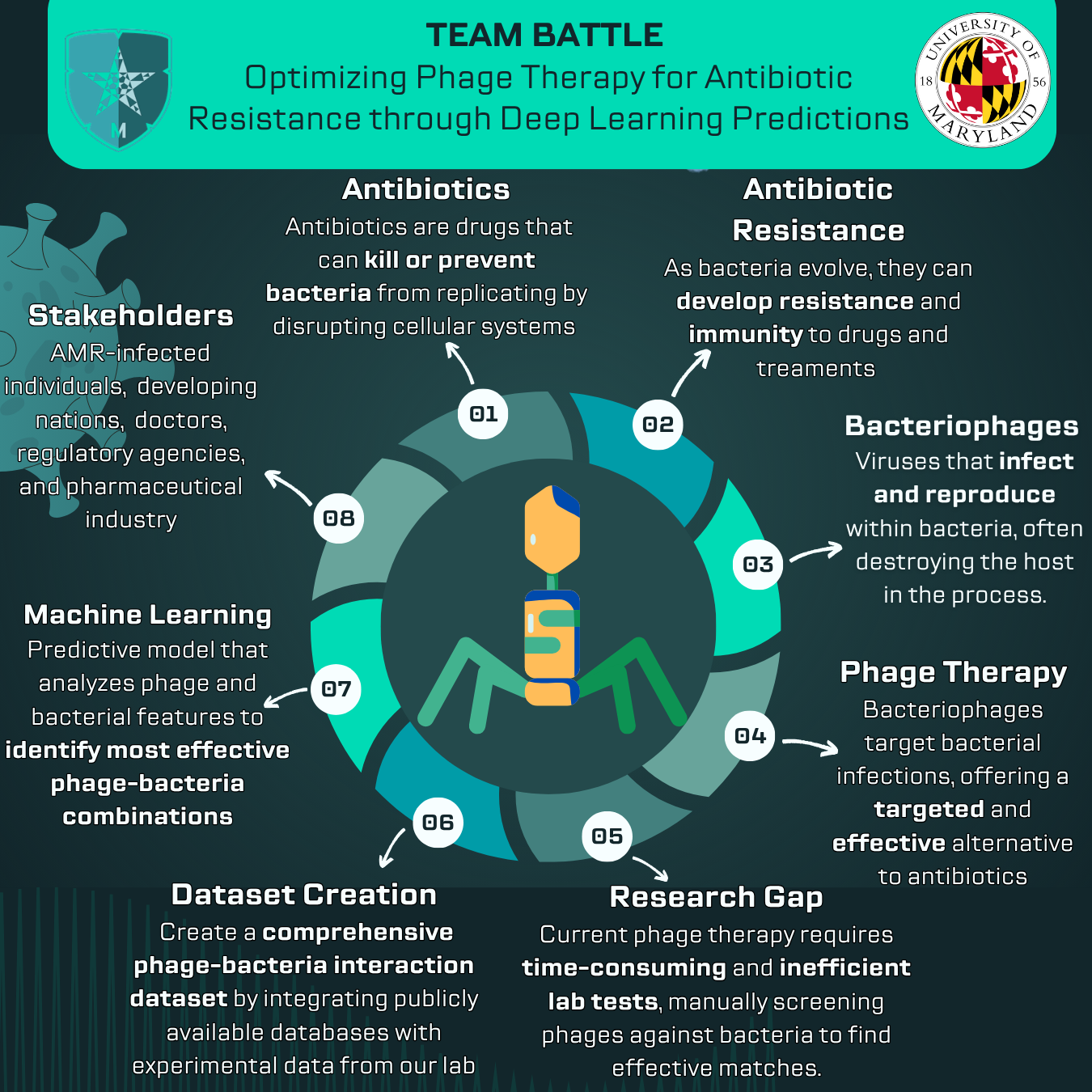
Antimicrobial resistance is a growing global health threat. As antibiotics lose effectiveness, there is urgent need for therapies that can target resistant bacteria with greater precision. Bacteriophages are promising because they naturally infect bacteria and can be selected for narrow, intentional targeting.
However, clinical use is still constrained by a major bottleneck: quickly identifying phages that will effectively lyse a specific patient isolate. This matching process is often labor intensive and time sensitive.
Investigate whether genome-scale language modeling can help generate viable candidate phages with lytic characteristics.
Develop models that predict infectivity likelihood using genomic and protein-level features from both phage and host.
Model infection dynamics over time to estimate treatment-relevant outcomes such as kill curves and resistance behavior.
Use plaque assays, titer measurements, and growth/lysis experiments to validate and continuously refine model predictions.