Project Progress
Key updates from our computational and experimental workstreams.
Update: model fitting for phage lysis challenge data
We fit a phage-host infection model to challenge data from S. aureus NRS382 with phage GRCS. In short, this compares predicted lysis dynamics against observed PFU and host trends to evaluate whether our model captures real infection behavior.
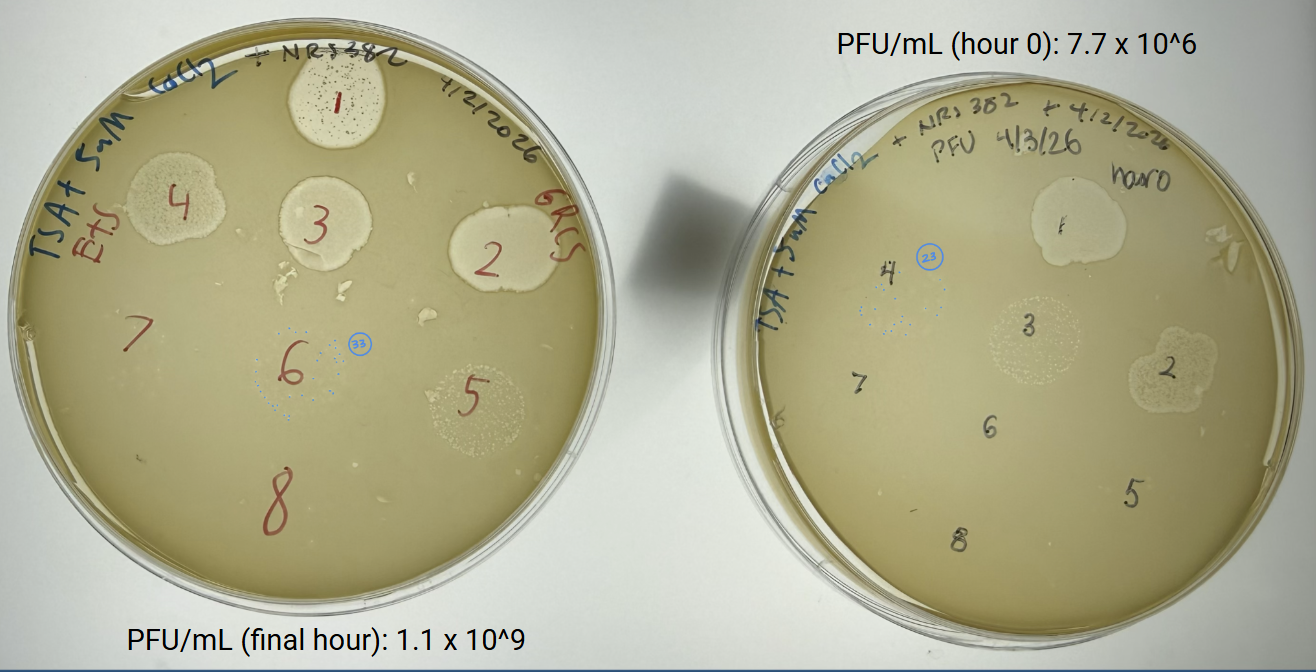

Experimental updates
- Completed growth-curve workflow for S. aureus JE2 with OD600 and CFU sampling.
- Ran phage lysis challenge with NRS382 and GRCS, including control comparison and titer measurements.
- Continued protocol development for titering, propagation, and serial dilution workflows.
Computational updates
- Joined and began onboarding on HPC resources for larger-scale modeling and parameter sweeps.
- Reproduced challenge systems inspired by latent-period inference work and one-step growth curve dynamics.
- Implemented ODE-based simulations across multiple phage-host systems to compare expected versus observed trajectories.
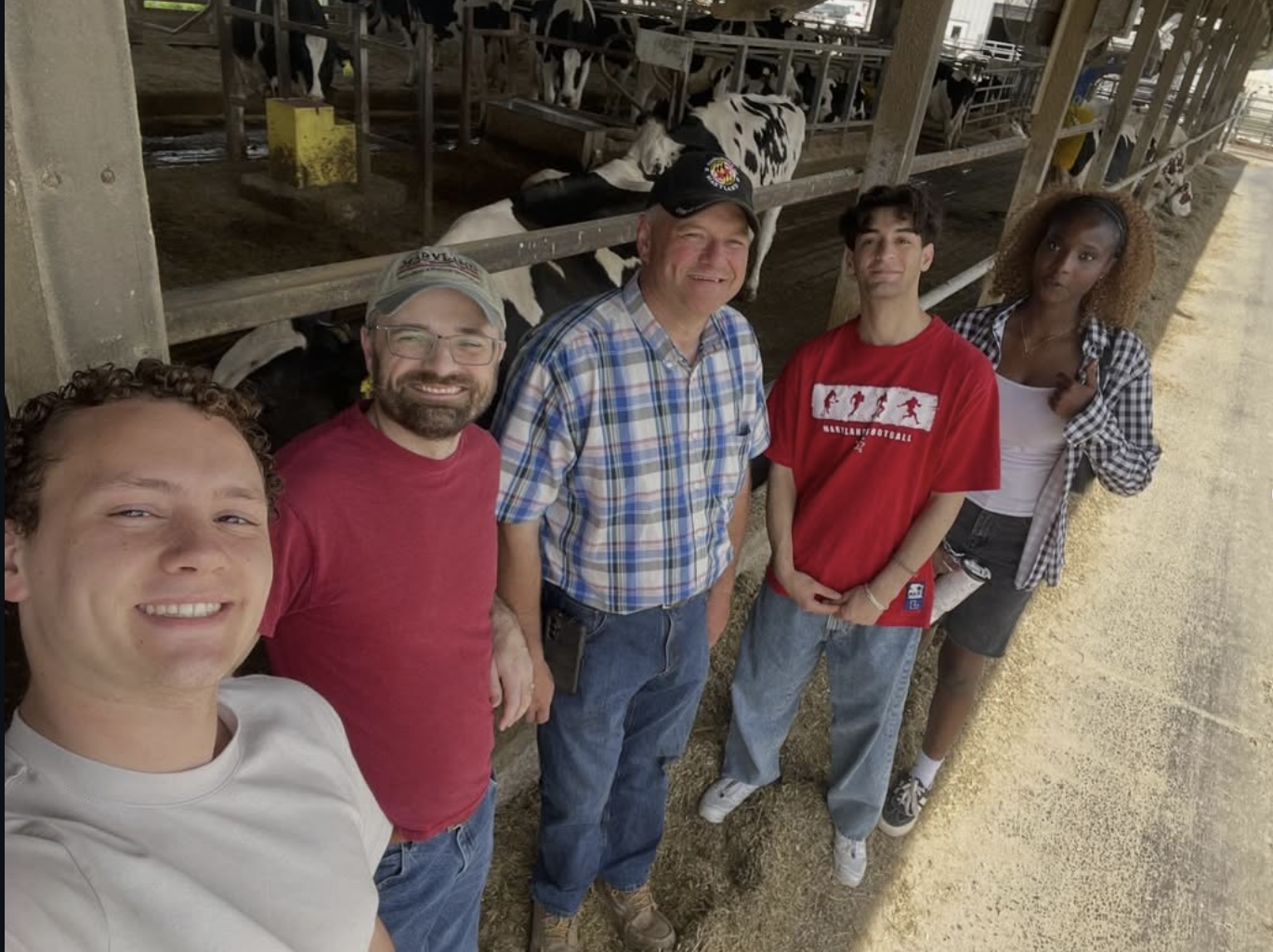
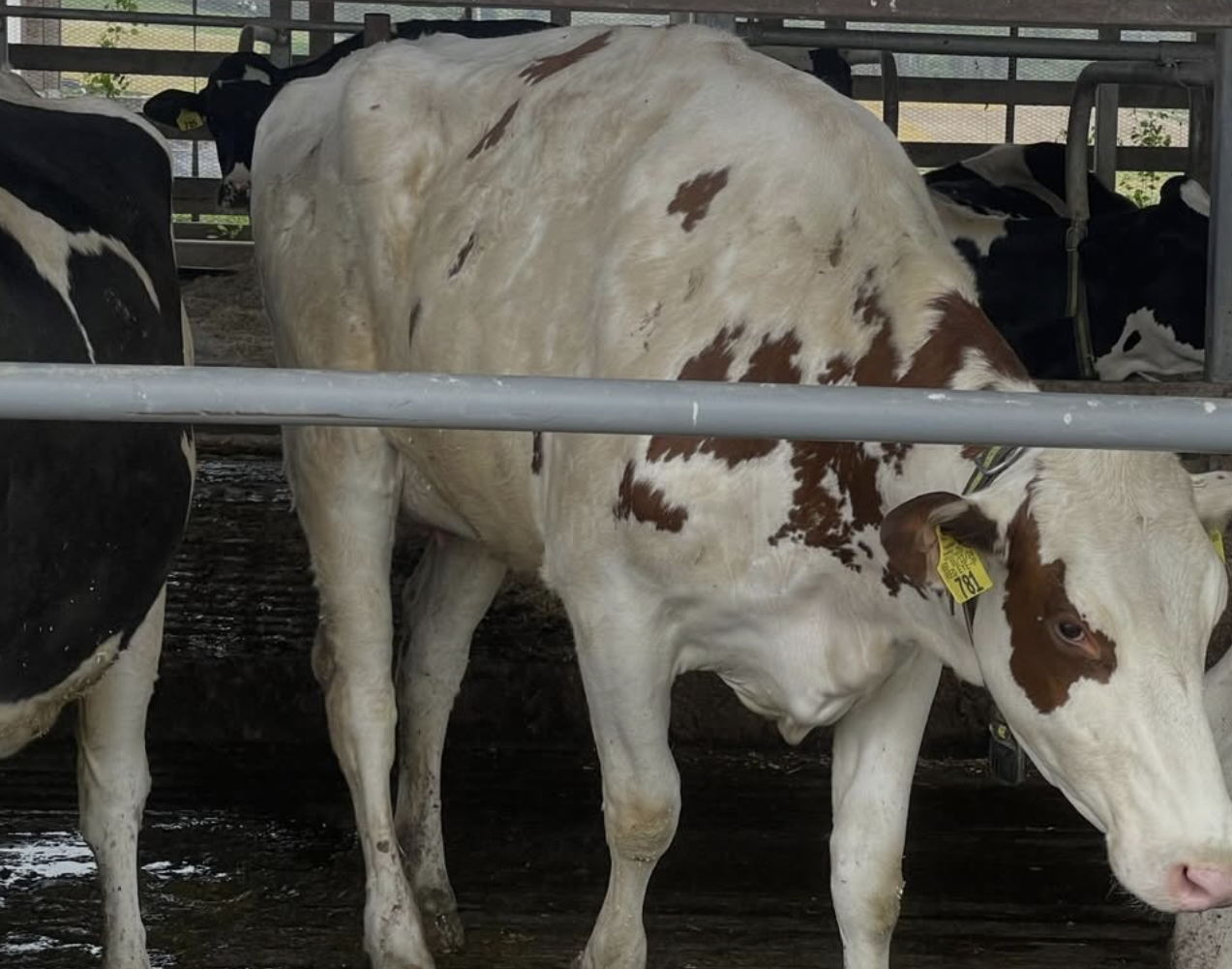
Field collaboration update: Central Maryland Research & Education Center
Our team visited the Central Maryland Research & Education Center with Brian S. Spielman (Assistant Director) to discuss how high somatic cell count dairy samples can support bacteriophage-focused infectious disease research. We observed the milking workflow and discussed sample collection logistics for downstream lab analysis.


Near-term next steps
- Expand host-strain panel for EoP comparisons and improved generalization checks.
- Strengthen calibration linking OD600 and CFU measurements for more stable joint fitting.
- Continue integrating experimental feedback to refine the scoring and simulation pipeline.